The Shanghai - Melbourne Research Training Group was founded in 2020 to develop the next generation of globally-engaged researchers. Researchers, together with joint PhD candidates, from both Shanghai Jiao Tong University (SJTU) and The University of Melbourne, work together on projects that aim to address the shared challenges of today and tomorrow, like climate change and global health.
Researchers involved in the Shanghai - Melbourne Research Training Group access the extensive resources and facilities both institutions have to offer. Candidates are jointly supervised by global experts from SJTU and Melbourne and become part of a growing network of academics with experience in the research landscape in both China and Australia.
Our partner: Shanghai Jiao Tong University
Founded in 1896, Shanghai Jiao Tong University (SJTU) is one of China’s oldest and most prestigious research universities. SJTU hosts many research institutes across the sciences and the humanities, leading innovation on the world stage. SJTU is one of the top research institutions in the country in terms of both the number of research projects and the amount of research funding received from the National Natural Science Foundation of China.

I had a great time studying from the best of both universities thanks to the support from the joint PhD team and my supervisors.Gabriel J. Gotama Shanghai Jiao Tong University - Melbourne joint PhD candidate
Project spotlight
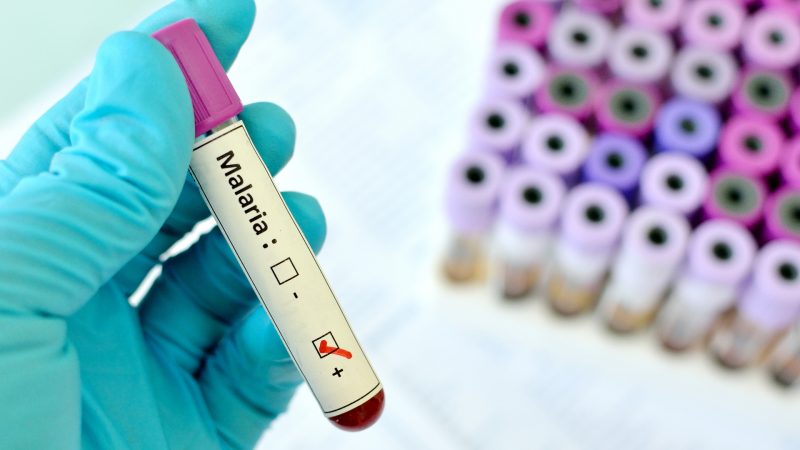
Learning from China’s successes to inform malaria elimination in the Asia-Pacific
China has not reported any local malaria cases since 2017 despite importations from neighbouring endemic countries. The project aims to identify the critical interventions to this success and integrate them into existing mathematical models of malaria transmission. These models will then be examined to see how similar interventions can help Asia-Pacific nations overcome the final barriers to malaria elimination. The project will combine in-depth epidemiological analyses with advanced mathematical models.
在联合博士团队和我的导师的支持下,我体验和感受了这两所顶尖大学所赋予的学术精髓。Gabriel J. Gotama 上海交通大学与墨尔本大学联合博士生
Meet our academic lead
Associate Professor Yi Yang is a faculty member of the Department of Mechanical Engineering at the University of Melbourne. Professor Yang’s research aims to develop next-generation engines with low greenhouse gas and pollutant emissions. His interests include alternative fuels, advanced compression ignition engines, fuel/engine interactions, combustion chemistry, renewable fuel production, catalytic combustion, and emission control.

First published on 30 August 2022.
Share this article
Related items
-
International Research Training Groups
IRTGs are cohort-based programs where graduate researchers enrol at both the University of Melbourne and an international partner university.
-
Find a supervisor or research project
Discover how to find a supervisor and learn how they can support your graduate research, or search available research projects.
-
Research degrees
Become a graduate researcher at the University of Melbourne.
-
Your study experience
Discover what it's like to be a graduate researcher. Find out about University life, support services, and opportunities for skills development.